Identifying the transcriptional regulatory sequences in genomes
With the rapid advances in sequencing technologies, obtaining the genome sequences of an individual organism is no longer rate limiting. Instead, identifying the functional elements throughout the genome has become a major bottleneck.
More ›
Epigenetic mechanisms regulating pluripotency and lineage commitment
We have generated comprehensive epigenome maps for the human embryonic stem cells (ESC), fibroblasts and a number of ES cell derived cell types. Analysis of these epigenomic profiles has revealed dramatic differences of DNA methylomes and chromatin landscapes between the pluripotent and lineage-committed cell types.
More ›
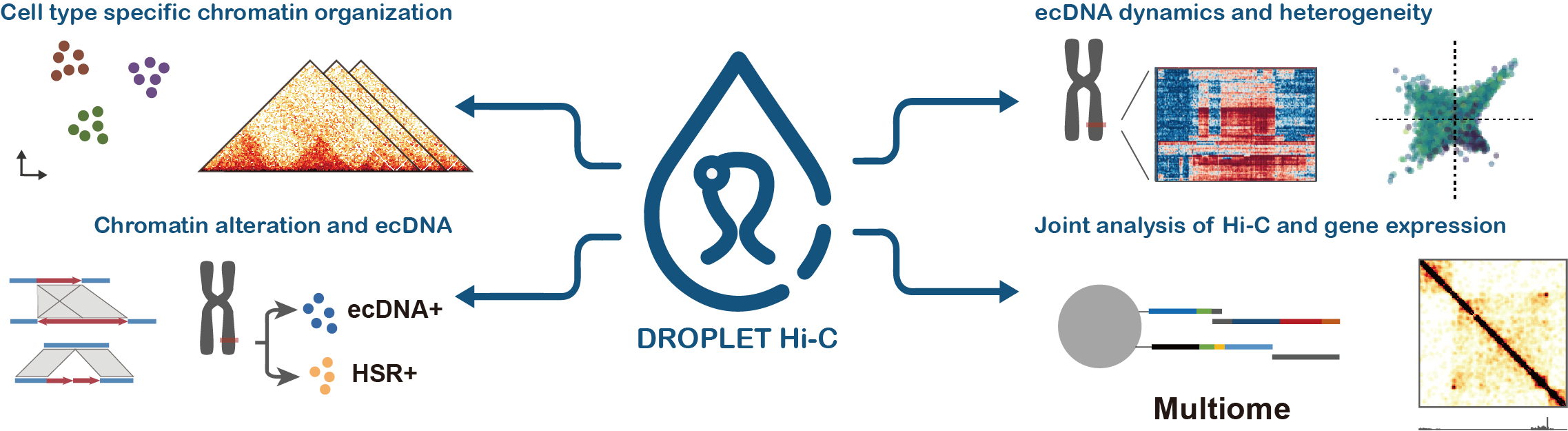
Higher-order genome architecture
Higher-order chromatin architecture is emerging as an important regulator of diverse nuclear processes, from gene regulation to DNA replication. Recent methodological advancements have allowed, for the first time, the ability to interrogate higher-order chromatin interactions on a genome-wide scale.
More ›